Biopython version 1.87
© 2026. All rights reserved.
© 2026. All rights reserved.
Bio.PDB is a Biopython module that focuses on working with crystal
structures of biological macromolecules. This document gives a fairly
complete overview of Bio.PDB.
Bio.PDB is automatically installed as part of Biopython. Biopython can
be obtained from http://www.biopython.org. It runs on many platforms
(Linux/Unix, windows, Mac,…).
Bio.PDB was used in the construction of DISEMBL, a web
server that predicts disordered regions in proteins , and COLUMBA, a website
that provides annotated protein structures (not longer available?). Bio.PDB
has also been used to perform a large scale search for active sites similarities
between protein structures in the PDB (see Proteins 51: 96–108,
2003), and to develop a new
algorithm that identifies linear secondary structure elements (see BMC
Bioinformatics 6: 202,
2005).
Judging from requests for features and information, Bio.PDB is also used
by several LPCs (Large Pharmaceutical Companies :-).
Yes, and I’d appreciate it if you would refer to Bio.PDB in publications
if you make use of it. The reference is:
Hamelryck, T., Manderick, B. (2003) PDB parser and structure class implemented in Python. Bioinformatics 19: 2308–2310
The article can be freely downloaded via the Bioinformatics journal
website. I
welcome e-mails telling me what you are using Bio.PDB for. Feature
requests are welcome too.
Pretty well, actually. Bio.PDB has been extensively tested on nearly
5500 structures from the PDB - all structures seemed to be parsed
correctly. More details can be found in the Bio.PDB Bioinformatics
article. Bio.PDB has been used/is being used in many research projects
as a reliable tool. In fact, I’m using Bio.PDB almost daily for research
purposes and continue working on improving it and adding new features.
The PDBParser performance was tested on about 800 structures (each
belonging to a unique SCOP superfamily). This takes about 20 minutes, or
on average 1.5 seconds per structure. Parsing the structure of the large
ribosomal subunit (1FKK), which contains about 64000 atoms, takes 10
seconds on a 1000 MHz PC. In short: it’s more than fast enough for many
applications.
Bio.PDB might be exactly what you want, and then again it might not. If
you are interested in data mining the PDB header, you might want to look
elsewhere because there is only limited support for this. If you look
for a powerful, complete data structure to access the atomic data
Bio.PDB is probably for you.
That’s simple:
from Bio.PDB import *
Not directly, mostly since there are quite a few Python based/Python
aware solutions already, that can potentially be used with Bio.PDB. My
choice is Pymol, BTW (I’ve used this successfully with Bio.PDB, and
there will probably be specific PyMol modules in Bio.PDB soon/some day).
Python based/aware molecular graphics solutions include:
I’d be crazy to write another molecular graphics application (been there done that, actually :-).
You can however interactively view Biopython structural entities in a Jupyter notebook using nglview:
import nglview as nv
view = nv.show_biopython(structure)
view
First, create a PDBParser object:
parser = PDBParser()
Then, create a structure object from a PDB file in the following way
(the PDB file in this case is called 1FAT.pdb, PHA-L is a user
defined name for the structure):
structure = parser.get_structure("PHA-L", "1FAT.pdb")
Similarly to the case the case of PDB files, first create an
MMCIFParser object:
parser = MMCIFParser()
Then use this parser to create a structure object from the mmCIF file:
structure = parser.get_structure("PHA-L", "1FAT.cif")
That’s not yet supported, but I’m definitely planning to support that in the future (it’s not a lot of work). Contact me if you need this, it might encourage me :-).
You got it. You can create a Python dictionary that maps all mmCIF tags
in an mmCIF file to their values. If there are multiple values (like in
the case of tag _atom_site.Cartn_y, which holds the y coordinates of
all atoms), the tag is mapped to a list of values. The dictionary is
created from the mmCIF file as follows:
from Bio.PDB.MMCIF2Dict import MMCIF2Dict
mmcif_dict = MMCIF2Dict("1FAT.cif")
Example: get the solvent content from an mmCIF file:
sc = mmcif_dict["_exptl_crystal.density_percent_sol"]
Example: get the list of the y coordinates of all atoms
y_list = mmcif_dict["_atom_site.Cartn_y"]
Thanks to Christian Rother you can access some information from the PDB
header. Note however that many PDB files contain headers with incomplete
or erroneous information. Many of the errors have been fixed in the
equivalent mmCIF files. Hence, if you are interested in the header
information, it is a good idea to extract information from mmCIF files
using the MMCIF2Dict tool described above, instead of parsing the PDB
header.
Now that is clarified, let’s return to parsing the PDB header. The
structure object has an attribute called header which is a python
dictionary that maps header records to their values.
Example:
resolution = structure.header["resolution"]
keywords = structure.header["keywords"]
The available keys are name, head, deposition_date,
release_date, structure_method, resolution, structure_reference
(maps to a list of references), journal_reference, author and
compound (maps to a dictionary with various information about the
crystallized compound).
The dictionary can also be created without creating a Structure
object, ie. directly from the PDB file:
handle = open(filename, "r")
header_dict = parse_pdb_header(handle)
handle.close()
Sure. Many PDB parsers assume that there is only one model, making them
all but useless for NMR structures. The design of the Structure object
makes it easy to handle PDB files with more than one model (see section
The Structure Object).
This can be done using the PDBList object, using the
retrieve_pdb_file method. The argument for this method is the PDB
identifier of the structure.
pdbl = PDBList()
pdbl.retrieve_pdb_file("1FAT")
The PDBList class can also be used as a command-line tool:
python PDBList.py 1fat
The downloaded file will be called pdb1fat.ent and
stored in the current working directory. Note that the
retrieve_pdb_file method also has an optional argument pdir that
specifies a specific directory in which to store the downloaded PDB
files.
The retrieve_pdb_file method also has some options for specifying the
compression format used for the download, and the program used for local
decompression (default .Z format and gunzip). In addition, the PDB
ftp site can be specified upon creation of the PDBList object. By
default, the ftp server of the Worldwide Protein Data
Bank is used. See
the API documentation for more details. Thanks again to Kristian Rother
for donating this module.
The following commands will store all PDB files in the /data/pdb
directory:
python PDBList.py all /data/pdb
python PDBList.py all /data/pdb -d
The API method for this is called download_entire_pdb. Adding the -d
option will store all files in the same directory. Otherwise, they are
sorted into PDB-style subdirectories according to their PDB ID’s.
Depending on the traffic, a complete download will take 2-4 days.
This can also be done using the PDBList object. One simply creates a
PDBList object (specifying the directory where the local copy of the
PDB is present) and calls the update_pdb method:
pl = PDBList(pdb="/data/pdb")
pl.update_pdb()
One can of course make a weekly cronjob out of this to keep the local copy automatically up-to-date. The PDB ftp site can also be specified (see API documentation).
PDBList has some additional methods that can be of use. The
get_all_obsolete method can be used to get a list of all obsolete PDB
entries. The changed_this_week method can be used to obtain the
entries that were added, modified or obsoleted during the current week.
For more info on the possibilities of PDBList, see the API
documentation.
It is well known that many PDB files contain semantic errors (I’m not
talking about the structures themselves, but their representation
in PDB files). Bio.PDB tries to handle this in two ways. The PDBParser
object can behave in two ways: a restrictive way and a permissive way
(THIS IS NOW THE DEFAULT). The restrictive way used to be the default,
but people seemed to think that Bio.PDB ‘crashed’ due to a bug (hah!),
so I changed it. If you ever encounter a real bug, please tell me
immediately!
Example:
# Permissive parser
parser = PDBParser(PERMISSIVE=1)
parser = PDBParser() # The same (default)
# Strict parser
strict_parser = PDBParser(PERMISSIVE=0)
In the permissive state (DEFAULT), PDB files that obviously contain errors are ‘corrected’ (i.e. some residues or atoms are left out). These errors include:
These errors indicate real problems in the PDB file (for details see the Bioinformatics article). In the restrictive state, PDB files with errors cause an exception to occur. This is useful to find errors in PDB files.
Some errors however are automatically corrected. Normally each disordered atom should have a non-blank altloc identifier. However, there are many structures that do not follow this convention, and have a blank and a non-blank identifier for two disordered positions of the same atom. This is automatically interpreted in the right way.
Sometimes a structure contains a list of residues belonging to chain A, followed by residues belonging to chain B, and again followed by residues belonging to chain A, i.e. the chains are ‘broken’. This is also correctly interpreted.
Use the PDBIO class for this. It’s easy to write out specific parts of a
structure too, of course.
Example: saving a structure
io = PDBIO()
io.set_structure(s)
io.save("out.pdb")
If you want to write out a part of the structure, make use of the
Select class (also in PDBIO). Select has four methods:
accept_model(model)
accept_chain(chain)
accept_residue(residue)
accept_atom(atom)
By default, every method returns 1 (which means the
model/chain/residue/atom is included in the output). By subclassing
Select and returning 0 when appropriate you can exclude models,
chains, etc. from the output. Cumbersome maybe, but very powerful. The
following code only writes out glycine residues:
class GlySelect(Select):
def accept_residue(self, residue):
if residue.get_name() == "GLY":
return 1
else:
return 0
io = PDBIO()
io.set_structure(s)
io.save("gly_only.pdb", GlySelect())
If this is all too complicated for you, the Dice module contains a
handy extract function that writes out all residues in a chain between
a start and end residue.
No, and I also don’t have plans to add that functionality soon (or ever—I don’t need it at all, and it’s a lot of work, plus no-one has ever asked for it). People who want to add this can contact me.
The Structure object follows the so-called SMCRA
(Structure/Model/Chain/Residue/Atom) architecture :
This is the way many structural biologists/bioinformaticians think about
structure, and provides a simple but efficient way to deal with
structure. Additional stuff is essentially added when needed. A UML
diagram of the Structure object (forgetting about the Disordered
classes for now) is shown in the figure below.
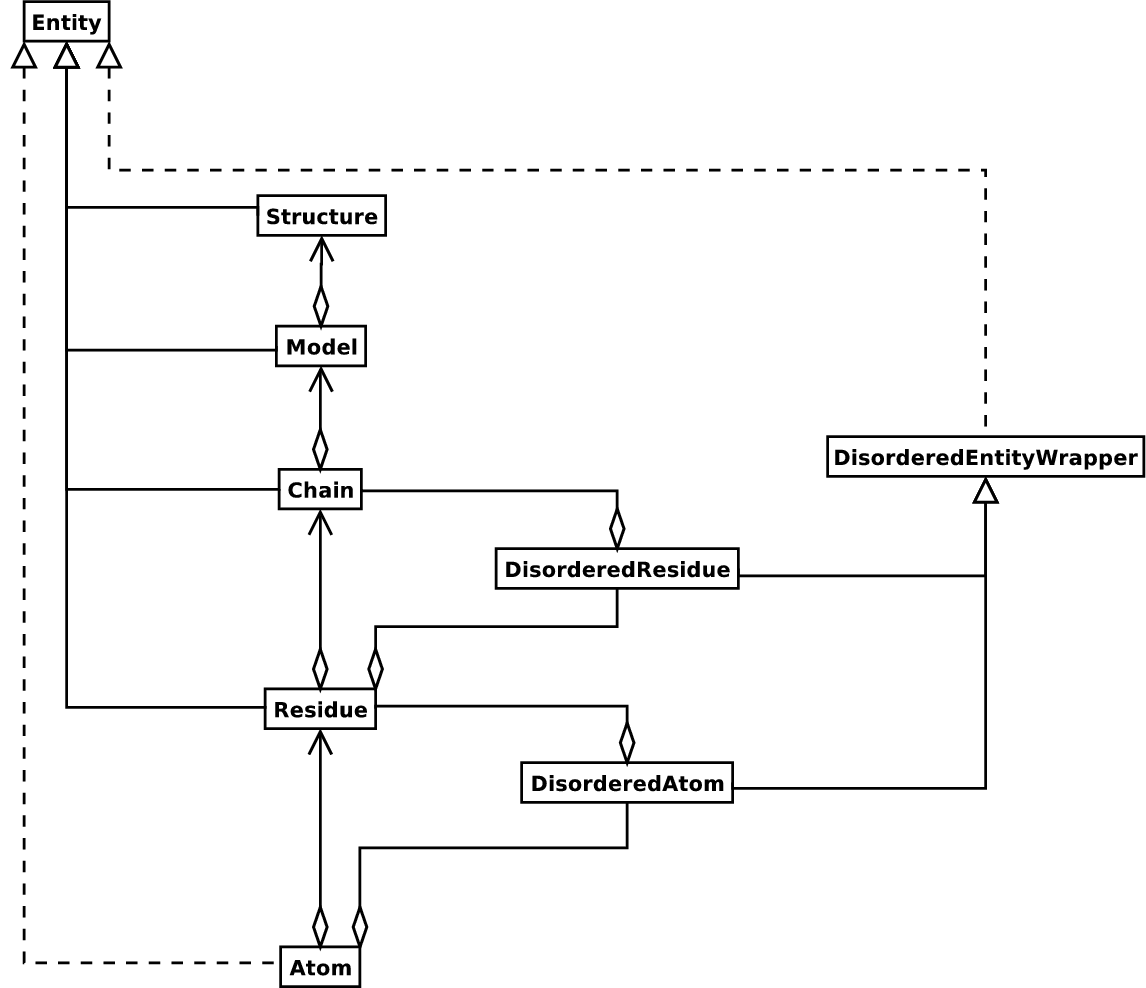
Diagram of SMCRA architecture of the Structure object.
Full lines with diamonds denote aggregation, full lines with arrows denote referencing, full lines with triangles denote inheritance and dashed lines with triangles denote interface realization.
The following code iterates through all atoms of a structure:
p = PDBParser()
structure = p.get_structure("X", "pdb1fat.ent")
for model in structure:
for chain in model:
for residue in chain:
for atom in residue:
print(atom)
There are also some shortcuts:
# Iterate over all atoms in a structure
for atom in structure.get_atoms():
print(atom)
# Iterate over all residues in a model
for residue in model.get_residues():
print(residue)
Structures, models, chains, residues and atoms are called Entities
in Biopython. You can always get a parent Entity from a child
Entity, e.g.:
residue = atom.get_parent()
chain = residue.get_parent()
You can also test whether an Entity has a certain child using the
has_id method.
You can do things like:
atoms = structure.get_atoms()
residues = structure.get_residues()
atoms = chain.get_atoms()
You can also use the Selection.unfold_entities function:
# Get all residues from a structure
res_list = Selection.unfold_entities(structure, "R")
# Get all atoms from a chain
atom_list = Selection.unfold_entities(chain, "A")
Obviously, A=atom, R=residue, C=chain, M=model, S=structure.
You can use this to go up in the hierarchy, e.g. to get a list of
(unique) Residue or Chain parents from a list of Atoms:
residue_list = Selection.unfold_entities(atom_list, "R")
chain_list = Selection.unfold_entities(atom_list, "C")
For more info, see the API documentation.
Atom/Residue/Chain/Model from a Structure?Easy. Here are some examples:
model = structure[0]
chain = model["A"]
residue = chain[100]
atom = residue["CA"]
Note that you can use a shortcut:
atom = structure[0]["A"][100]["CA"]
The model id is an integer which denotes the rank of the model in the PDB/mmCIF file. The model id starts at 0. Crystal structures generally have only one model (with id 0), while NMR files usually have several models.
The chain id is specified in the PDB/mmCIF file, and is a single character (typically a letter).
This is a bit more complicated, due to the clumsy PDB format. A residue id is a tuple with three elements:
'H_' plus the name of the
hetero-residue (e.g. 'H_GLC' in the case of a glucose molecule),
or 'W' in the case of a water molecule.'A'. The insertion code is sometimes
used to preserve a certain desirable residue numbering scheme. A Ser
80 insertion mutant (inserted e.g. between a Thr 80 and an Asn
81 residue) could e.g. have sequence identifiers and insertion codes
as follows: Thr 80 A, Ser 80 B, Asn 81. In this way the residue
numbering scheme stays in tune with that of the wild type structure.The id of the above glucose residue would thus be ('H_GLC', 100, 'A').
If the hetero-flag and insertion code are blank, the sequence
identifier alone can be used:
# Full id
residue = chain[(" ", 100, " ")]
# Shortcut id
residue = chain[100]
The reason for the hetero-flag is that many, many PDB files use the same sequence identifier for an amino acid and a hetero-residue or a water, which would create obvious problems if the hetero-flag was not used.
The atom id is simply the atom name (eg. 'CA'). In practice, the atom
name is created by stripping all spaces from the atom name in the PDB
file.
However, in PDB files, a space can be part of an atom name. Often,
calcium atoms are called 'CA..' in order to distinguish them from Cα
atoms (which are called '.CA.'). In cases were stripping the spaces
would create problems (ie. two atoms called 'CA' in the same residue)
the spaces are kept.
This is one of the strong points of Bio.PDB. It can handle both
disordered atoms and point mutations (ie. a Gly and an Ala residue in
the same position).
Disorder should be dealt with from two points of view: the atom and the residue points of view. In general, I have tried to encapsulate all the complexity that arises from disorder. If you just want to loop over all Cα atoms, you do not care that some residues have a disordered side chain. On the other hand it should also be possible to represent disorder completely in the data structure. Therefore, disordered atoms or residues are stored in special objects that behave as if there is no disorder. This is done by only representing a subset of the disordered atoms or residues. Which subset is picked (e.g. which of the two disordered OG side chain atom positions of a Ser residue is used) can be specified by the user.
Disordered atom positions are represented by ordinary Atom
objects, but all Atom objects that represent the same physical atom
are stored in a DisorderedAtom object (see section The Structure
Object). Each Atom object in a
DisorderedAtom object can be uniquely indexed using its altloc
specifier. The DisorderedAtom object forwards all uncaught method
calls to the selected Atom object, by default the one that represents
the atom with the highest occupancy. The user can of course change the
selected Atom object, making use of its altloc specifier. In this way
atom disorder is represented correctly without much additional
complexity. In other words, if you are not interested in atom disorder,
you will not be bothered by it.
Each disordered atom has a characteristic altloc identifier. You can
specify that a DisorderedAtom object should behave like the Atom
object associated with a specific altloc identifier:
atom.disordered_select("A") # select altloc A atom
atom.disordered_select("B") # select altloc B atom
A special case arises when disorder is due to point mutations, i.e. when two or more point mutants of a polypeptide are present in the crystal. An example of this can be found in PDB structure 1EN2.
Since these residues belong to a different residue type (e.g. let’s say
Ser 60 and Cys 60) they should not be stored in a single Residue
object as in the common case. In this case, each residue is represented
by one Residue object, and both Residue objects are stored in a
single DisorderedResidue object (see section The Structure
Object).
The DisorderedResidue object forwards all uncaught methods to the
selected Residue object (by default the last Residue object added),
and thus behaves like an ordinary residue. Each Residue object in a
DisorderedResidue object can be uniquely identified by its residue
name. In the above example, residue Ser 60 would have id ‘SER’ in the
DisorderedResidue object, while residue Cys 60 would have id ‘CYS’.
The user can select the active Residue object in a DisorderedResidue
object via this id.
Example: suppose that a chain has a point mutation at position 10, consisting of a Ser and a Cys residue. Make sure that residue 10 of this chain behaves as the Cys residue.
residue = chain[10]
residue.disordered_select("CYS")
In addition, you can get a list of all Atom objects (ie. all
DisorderedAtom objects are ‘unpacked’ to their individual Atom
objects) using the get_unpacked_list method of a
(Disordered)Residue object.
Yes, kinda, but I’m waiting for a request for this feature to finish it :-).
Well, yes! Bio.PDB supports isotropic and anisotropic B factors, and
also deals with standard deviations of anisotropic B factor if present
(see the section Analysis).
Yup, supported. See the section Analysis.
Sure, sure. Everybody is always coming up with (mostly vaporware or
partly implemented) data structures that handle all possible situations
and are extensible in all thinkable (and unthinkable) ways. The prosaic
truth however is that 99.9% of people using (and I mean really using!)
crystal structures think in terms of models, chains, residues and atoms.
The philosophy of Bio.PDB is to provide a reasonably fast, clean,
simple, but complete data structure to access structure data. The proof
of the pudding is in the eating.
Moreover, it is quite easy to build more specialised data structures on
top of the Structure class (eg. there’s a Polypeptide class). On the
other hand, the Structure object is built using a Parser/Consumer
approach (called PDBParser/MMCIFParser and StructureBuilder,
respectively). One can easily reuse the PDB/mmCIF parsers by
implementing a specialised StructureBuilder class. It is of course
also trivial to add support for new file formats by writing new parsers.
Atom object?Using the following methods:
a.get_name() # atom name (spaces stripped, e.g. 'CA')
a.get_id() # id (equals atom name)
a.get_coord() # atomic coordinates
a.get_vector() # atomic coordinates as Vector object
a.get_bfactor() # isotropic B factor
a.get_occupancy() # occupancy
a.get_altloc() # alternative location specifier
a.get_sigatm() # std. dev. of atomic parameters
a.get_siguij() # std. dev. of anisotropic B factor
a.get_anisou() # anisotropic B factor
a.get_fullname() # atom name (with spaces, e.g. '.CA.')
Residue object?Using the following methods:
r.get_resname() # return the residue name (eg. 'GLY')
r.is_disordered() # 1 if the residue has disordered atoms
r.get_segid() # return the SEGID
r.has_id(name) # test if a residue has a certain atom
That’s simple: the minus operator for atoms has been overloaded to return the distance between two atoms.
Example:
# Get some atoms
ca1 = residue1["CA"]
ca2 = residue2["CA"]
# Simply subtract the atoms to get their distance
distance = ca1 - ca2
This can easily be done via the vector representation of the atomic
coordinates, and the calc_angle function from the Vector module:
vector1 = atom1.get_vector()
vector2 = atom2.get_vector()
vector3 = atom3.get_vector()
angle = calc_angle(vector1, vector2, vector3)
Again, this can easily be done via the vector representation of the
atomic coordinates, this time using the calc_dihedral function from
the Vector module:
vector1 = atom1.get_vector()
vector2 = atom2.get_vector()
vector3 = atom3.get_vector()
vector4 = atom4.get_vector()
angle = calc_dihedral(vector1, vector2, vector3, vector4)
Use NeighborSearch. This uses a KD tree data structure coded in C
behind the screens, so it’s pretty darn fast (see Bio.KDTree).
Structure object?Use PolypeptideBuilder. You can use the resulting Polypeptide object
to get the sequence as a Seq object or to get a list of Cα atoms as
well. Polypeptides can be built using a C-N or a Cα-Cα distance
criterion.
Example:
# Using C-N
ppb = PPBuilder()
for pp in ppb.build_peptides(structure):
print(pp.get_sequence())
# Using CA-CA
ppb = CaPPBuilder()
for pp in ppb.build_peptides(structure):
print(pp.get_sequence())
Note that in the above case only model 0 of the structure is considered
by PolypeptideBuilder. However, it is possible to use
PolypeptideBuilder to build Polypeptide objects from Model and
Chain objects as well.
The first thing to do is to extract all polypeptides from the structure
(see previous entry). The sequence of each polypeptide can then easily
be obtained from the Polypeptide objects. The sequence is represented
as a Biopython Seq object.
Example:
>>> seq = polypeptide.get_sequence()
>>> print(seq)
Seq('SNVVE...')
For this functionality, you need to install
DSSP (and obtain a license
for it - free for academic use).
Then use the DSSP class, which maps Residue objects to their
secondary structure (and accessible surface area). The DSSP codes are
listed in the Table below. Note that DSSP (the program, and thus by
consequence the class) cannot handle multiple models!
| DSSP Code | Secondary structure |
|---|---|
| H | α-helix |
| B | Isolated β-bridge residue |
| E | Strand |
| G | 3-10 helix |
| I | Π-helix |
| T | Turn |
| S | Bend |
| - | Other |
Use the DSSP class (see also previous entry). But see also next entry.
Residue depth is the average distance of a residue’s atoms from the
solvent accessible surface. It’s a fairly new and very powerful
parameterization of solvent accessibility. For this functionality, you
need to install Michel Sanner’s MSMS
program.
Then use the ResidueDepth class. This class behaves as a dictionary
which maps Residue objects to corresponding (residue depth, Cα depth)
tuples. The Cα depth is the distance of a residue’s Cα atom to the
solvent accessible surface.
Example:
model = structure[0]
rd = ResidueDepth(model, pdb_file)
residue_depth, ca_depth = rd[some_residue]
You can also get access to the molecular surface itself (via the
get_surface function), in the form of a NumPy array with the
surface points.
Half Sphere Exposure (HSE) is a new, 2D measure of solvent exposure. Basically, it counts the number of Cα atoms around a residue in the direction of its side chain, and in the opposite direction (within a radius of 13 Å). Despite its simplicity, it outperforms many other measures of solvent exposure. An article describing this novel 2D measure has been submitted.
HSE comes in two flavors: HSEα and HSEβ. The former only uses the Cα
atom positions, while the latter uses the Cα and Cβ atom positions. The
HSE measure is calculated by the HSExposure class, which can also
calculate the contact number. The latter class has methods which return
dictionaries that map a Residue object to its corresponding HSEα, HSEβ
and contact number values.
Example:
model = structure[0]
hse = HSExposure()
# Calculate HSEalpha
exp_ca = hse.calc_hs_exposure(model, option="CA3")
# Calculate HSEbeta
exp_cb = hse.calc_hs_exposure(model, option="CB")
# Calculate classical coordination number
exp_fs = hse.calc_fs_exposure(model)
# Print HSEalpha for a residue
print(exp_ca[some_residue])
First, create an alignment file in FASTA format, then use the
StructureAlignment class. This class can also be used for alignments
with more than two structures.
Use is_aa(residue).
Atom objects return a Vector object representation of the
coordinates with the get_vector method. Vector implements the full
set of 3D vector operations, matrix multiplication (left and right) and
some advanced rotation-related operations as well. See also next
question.
OK, I admit, this example is only present to show off the possibilities
of Bio.PDB’s Vector module (though this code is actually used in the
HSExposure module, which contains a novel way to parametrize residue
exposure - publication underway). Suppose that you would like to find
the position of a Gly residue’s Cβ atom, if it had one. How would you do
that? Well, rotating the N atom of the Gly residue along the Cα-C bond
over -120 degrees roughly puts it in the position of a virtual Cβ atom.
Here’s how to do it, making use of the rotaxis method (which can be
used to construct a rotation around a certain axis) of the Vector
module:
# get atom coordinates as vectors
n = residue["N"].get_vector()
c = residue["C"].get_vector()
ca = residue["CA"].get_vector()
# center at origin
n = n - ca
c = c - ca
# find rotation matrix that rotates n -120 degrees along the ca-c vector
rot = rotaxis(-pi * 120.0 / 180.0, c)
# apply rotation to ca-n vector
cb_at_origin = n.left_multiply(rot)
# put on top of ca atom
cb = cb_at_origin + ca
This example shows that it’s possible to do some quite nontrivial vector
operations on atomic data, which can be quite useful. In addition to all
the usual vector operations (cross (use **), and dot (use *)
product, angle, norm, etc.) and the above mentioned rotaxis
function, the Vector module also has methods to rotate (rotmat)
or reflect (refmat) one vector on top of another.
Surprisingly, this is done using the Superimposer object. This object
calculates the rotation and translation matrix that rotates two lists of
atoms on top of each other in such a way that their RMSD is minimized.
Of course, the two lists need to contain the same amount of atoms. The
Superimposer object can also apply the rotation/translation to a list
of atoms. The rotation and translation are stored as a tuple in the
rotran attribute of the Superimposer object (note that the rotation
is right multiplying!). The RMSD is stored in the rmsd attribute.
The algorithm used by Superimposer comes from Matrix
computations, 2nd ed. Golub, G. & Van Loan
(1989)
and makes use of singular value decomposition (this is implemented in the
general Bio.SVDSuperimposer module).
Example:
sup = Superimposer()
# Specify the atom lists
# 'fixed' and 'moving' are lists of Atom objects
# The moving atoms will be put on the fixed atoms
sup.set_atoms(fixed, moving)
# Print rotation/translation/rmsd
print(sup.rotran)
print(sup.rms)
# Apply rotation/translation to the moving atoms
sup.apply(moving)
Pretty easily. Use the active site atoms to calculate the rotation/translation matrices (see above), and apply these to the whole molecule.
Yes, using the transform method of the Atom object, or directly
using the set_coord method.
See the main Biopython tutorial.
No documentation available yet.
Woah! It’s late and I’m tired, and a glass of excellent Pedro Ximenez sherry is waiting for me. Just drop me a mail, and I’ll answer you in the morning (with a bit of luck…).
The main author/maintainer of Bio.PDB is:
Thomas Hamelryck Bioinformatics center Institute of Molecular Biology University of Copenhagen Universitetsparken 15, Bygning 10 DK-2100 København Ø Denmark
Kristian Rother donated code to interact with the PDB database, and to
parse the PDB header. Indraneel Majumdar sent in some bug reports and
assisted in coding the Polypeptide module. Many thanks to Brad
Chapman, Jeffrey Chang, Andrew Dalke and Iddo Friedberg for suggestions,
comments, help and/or biting criticism :-).
Yes, yes, yes! Just send an e-mail to Thomas Hamelryck or to the Biopython developers if you have something useful to contribute! Eternal fame awaits!