Biopython version 1.87
© 2026. All rights reserved.
© 2026. All rights reserved.
This module provides classes, functions and I/O support for working with phylogenetic trees.
For more complete documentation, see the Phylogenetics chapter of the
Biopython
Tutorial and the
Bio.Phylo API
pages
generated from the source code. The Phylo
cookbook page has more examples of how to
use this module, and the PhyloXML page describes
how to attach graphical cues and additional information to a tree.
This module is included in Biopython 1.54 and later. If you’re interested in testing newer additions to this code before the next official release, see SourceCode for instructions on getting a copy of the development branch.
To draw trees (optional), you’ll also need these packages:
to_networkx (and deprecated function draw_graphviz)to_networkx (and deprecated function draw_graphviz)The I/O and tree-manipulation functionality will work without them;
they’re imported on demand when the functions draw(), draw_graphviz()
and to_networkx() are called.
The Phylo module has also been successfully tested on Jython 2.5.1,
minus the Graphviz- and NetworkX-based functions. However, parsing
phyloXML files is noticeably slower because Jython uses a different
version of the underlying XML parsing library.
Wrappers for supported file formats are available from the top level of the module:
from Bio import Phylo
Like SeqIO and AlignIO, this module
provides four I/O functions: parse(), read(), write() and convert().
Each function accepts either a file name or an open file handle, so data can
be also loaded from compressed files, StringIO objects, and so on. If the
file name is passed as a string, the file is automatically closed when the function
finishes; otherwise, you’re responsible for closing the handle yourself.
The second argument to each function is the target format. Currently, the following formats are supported:
See the PhyloXML page for more examples of using
tree objects.
Incrementally parse each tree in the given file or handle, returning an
iterator of Tree objects (i.e. some subclass of the Bio.Phylo.BaseTree
Tree class, depending on the file format).
>>> trees = Phylo.parse("phyloxml_examples.xml", "phyloxml")
>>> for tree in trees:
... print(tree.name)
...
example from Prof. Joe Felsenstein's book "Inferring Phylogenies"
example from Prof. Joe Felsenstein's book "Inferring Phylogenies"
same example, with support of type "bootstrap"
same example, with species and sequence
same example, with gene duplication information and sequence relationships
similar example, with more detailed sequence data
network, node B is connected to TWO nodes: AB and C
...
If there’s only one tree, then the next() method on the resulting
generator will return it.
>>> tree = Phylo.parse('phyloxml_examples.xml', 'phyloxml').next()
>>> tree.name
'example from Prof. Joe Felsenstein\'s book "Inferring Phylogenies"'
Note that this doesn’t immediately reveal whether there are any
remaining trees – if you want to verify that, use read() instead.
Parse and return exactly one tree from the given file or handle. If the
file contains zero or multiple trees, a ValueError is raised. This is
useful if you know a file contains just one tree, to load that tree
object directly rather than through parse() and next(), and as a safety
check to ensure the input file does in fact contain exactly one
phylogenetic tree at the top level.
tree = Phylo.read("example.dnd", "newick")
print(tree)
If you have your tree data already loaded as a Python string, you can
parse it with the help of StringIO (in Python’s standard library):
from cStringIO import StringIO
treedata = "(A, (B, C), (D, E))"
handle = StringIO(treedata)
tree = Phylo.read(handle, "newick")
In one line:
tree = Phylo.read(StringIO("(A, (B, C), (D, E))"), "newick")
The other I/O functions also can be used with StringIO.
(General tip: if you write to the StringIO object and want to re-read
the contents, you’ll need to call the seek(0) method to move the
handle back to the start of the StringIO data – the same as an open
file handle. See examples of this in the unit tests for
Phylo
in the Biopython source code.
Write a sequence of Tree objects to the given file or handle. Passing a
single Tree object instead of a list or iterable will also work (see,
Phylo is friendly).
tree1 = Phylo.read("example1.xml", "phyloxml")
tree2 = Phylo.read("example2.xml", "phyloxml")
Phylo.write([tree1, tree2], "example-both.xml", "phyloxml")
Given two files (or handles) and two formats, both supported by
Bio.Phylo, convert the first file from the first format to the second
format, writing the output to the second file.
Phylo.convert("example.nhx", "newick", "example2.nex", "nexus")
Within the Phylo module are parsers and writers for specific file
formats, conforming to the basic top-level API and sometimes adding
additional features.
PhyloXMLIO: Support for the phyloXML
format. See the PhyloXML page for details.
NeXMLIO: Support for the NeXML format.
NewickIO: A port of the parser in Bio.Nexus.Trees to support the
Newick (a.k.a. New Hampshire) format through the Phylo API.
NexusIO: Wrappers around Bio.Nexus to support the Nexus tree format.
CDAOIO: Support for the Comparative Data Analysis Ontology (CDAO). Requires RDFlib.
The Nexus format actually contains several sub-formats for different
kinds of data; to represent trees, Nexus provides a block containing
some metadata and one or more Newick trees (another kind of Nexus block
can represent alignments; this is handled in
AlignIO. So to parse a complete Nexus file with
all block types handled, use Bio.Nexus directly, and to extract just the
trees, use Bio.Phylo.
The basic objects are defined in Bio.Phylo.BaseTree.
To support additional information stored in specific file formats,
sub-modules within Tree offer additional classes that inherit from
BaseTree classes.
Each sub-class of BaseTree.Tree or Node has a class method to promote an
object from the basic type to the format-specific one. These sub-class
objects can generally be treated as instances of the basic type without
any explicit conversion.
PhyloXML: Support for the phyloXML format. See the
PhyloXML page for details.
Newick: The Newick module provides minor enhancements to the
BaseTree classes, plus several shims for compatibility with the existing
Bio.Nexus module. The API for this module is under development and
should not be relied on, other than the functionality already provided
by BaseTree.
Some additional tools are located in the Utils module under Bio.Phylo.
These functions are also loaded to the top level of the Phylo module on
import for easy access.
Where a third-party package is required, that package is imported when the function itself is called, so these dependencies are not necessary to install or use the rest of the Tree module.
str(tree) produces a plain-text representation of the entire
tree. Strings are automatically truncated to ensure reasonable display.
Use this with the print function to get a quick overview of your tree:
>>> tree = Phylo.parse('phyloxml_examples.xml', 'phyloxml').next()
>>> print(tree)
Phylogeny(description='phyloXML allows to use either a "branch_length"
attribute or element to indicate branch lengths.', name='example from
Prof. Joe Felsenstein s book "Inferring Phylogenies"')
Clade()
Clade(branch_length=0.06)
Clade(branch_length=0.102, name='A')
Clade(branch_length=0.23, name='B')
Clade(branch_length=0.4, name='C')
...
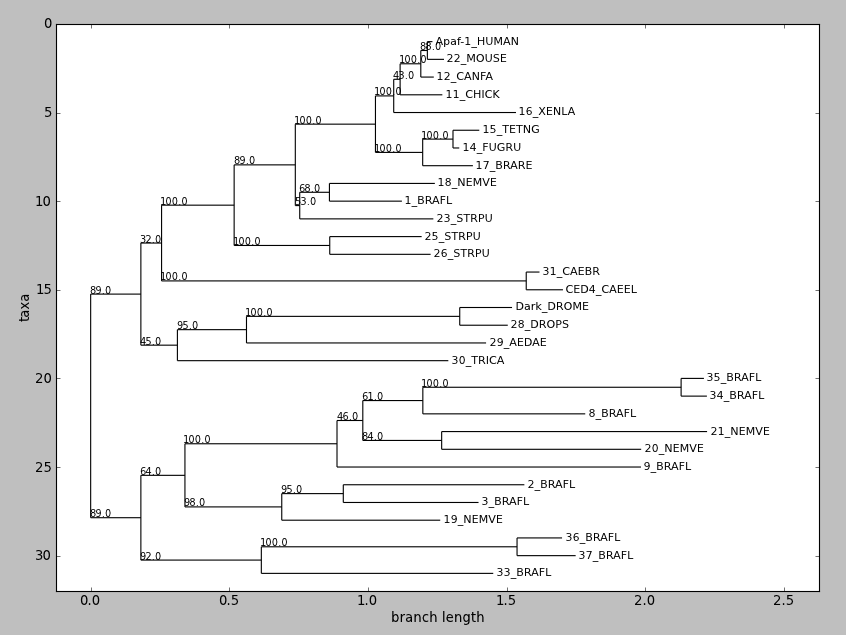
draw() displays a rooted phylogram using Matplotlib or Pylab. New in
Biopython 1.58.
Try this:
tree = Phylo.read("apaf.xml", "phyloxml")
tree.ladderize() # Flip branches so deeper clades are displayed at top
Phylo.draw(tree)
draw_graphviz mimics the networkx function of the same name, with
some tweaks to improve the display of the graph. If a file name is
given, the graph is drawn directly to that file, and options such as
image format (default PDF) may be used.
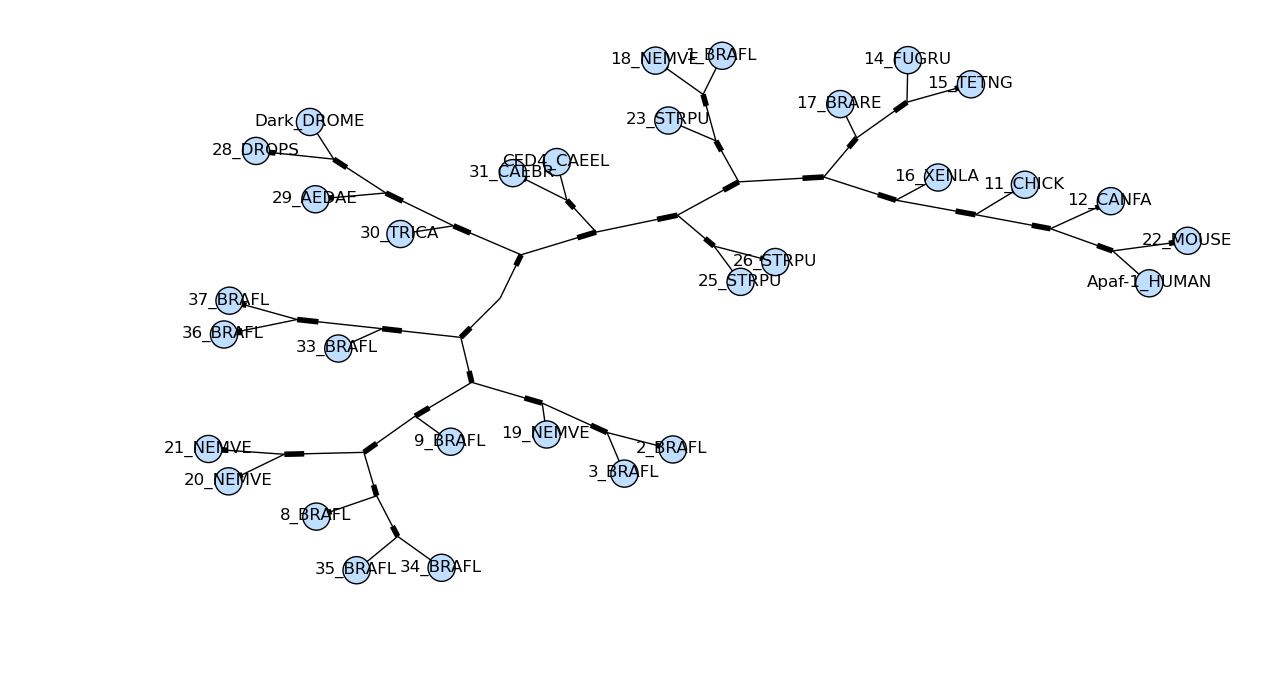
Sadly the plot draw_graphviz draws is misleading, so we have deprecated
this method. The branch lengths are ignored and the distances between nodes
in the plot is arbitrary, per the graphviz layout engine. But it looks like
a proper radial phylogeny at first glance, which could lead to a wrong
interpretation of the data. It would be better for users to create radial
plots with another library like ETE or DendroPy, or just use the simple
rectangular plot produced by Phylo.draw.
Prerequisites: In addition to NetworkX, you’ll need a local installation of Graphviz, Matplotlib and either PyGraphviz or pydot.
Drawing a basic dendrogram is simple:
import pylab
tree = Phylo.read("apaf.xml", "phyloxml")
Phylo.draw_graphviz(tree)
pylab.show()

Here’s the same tree without the circles at each labelled node:
Phylo.draw_graphviz(tree, node_size=0)
See the Phylo cookbook page for more drawing features and options.
draw_ascii prints an ascii-art rooted phylogram to standard output,
or another file handle if specified. Only terminal node labels are
shown; these are the result of str(clade) (usually clade names). The
width of the text field used for drawing is 80 characters by default,
adjustable with the column_width keyword argument, and the height in
character rows is twice the number of terminals in the tree.
A simple tree with defined branch lengths looks like this:
>>> tree = Phylo.parse('phyloxml_examples.xml', 'phyloxml').next()
>>> Phylo.draw_ascii(tree)
_____________ A
_______|
_| |_______________________________ B
|
|_______________________________________________________ C
The same topology without branch lengths is drawn with equal-length branches:
___________________________ A
___________________________|
_| |___________________________ B
|
|___________________________ C
A larger tree (apaf.xml, 31 leaf nodes) drawn with the default column width demonstrates how relatively short branches are handled:
>>> apaf = Phylo.read('apaf.xml', 'phyloxml')
>>> Phylo.draw_ascii(apaf)
_ 22_MOUSE
|
_| Apaf-1_HUMAN
| |
,| | 12_CANFA
||
_||___ 11_CHICK
| |
| |___________ 16_XENLA
_______|
| | , 14_FUGRU
| | __|
| |____| |__ 15_TETNG
_____| |
| | |____ 17_BRARE
| |
| | ______ 1_BRAFL
| | __|
______| || |_________ 18_NEMVE
| | |
| | |____________ 23_STRPU
| |
_| | _________ 26_STRPU
| | |_________|
| | |________ 25_STRPU
| |
| | ___ CED4_CAEEL
| |___________________________________|
____| |_ 31_CAEBR
| |
| | ___ 28_DROPS
| | _____________________|
| | ______| |____ Dark_DROME
| | | |
| |__| |_______________________ 29_AEDAE
| |
| |__________________________ 30_TRICA
|
| _ 34_BRAFL
| _________________________|
_| _____| |_ 35_BRAFL
| | |
| __| |_______________ 8_BRAFL
| | |
| | | ___________________ 20_NEMVE
| ______________| |_______|
| | | |__________________________ 21_NEMVE
| | |
| ___| |______________________________ 9_BRAFL
| | |
| | | _____________ 3_BRAFL
| | | _____|
| | |_________| |_________________ 2_BRAFL
|____| |
| |_______________ 19_NEMVE
|
| _____ 37_BRAFL
| ________________________|
|___________| |____ 36_BRAFL
|
|______________________ 33_BRAFL
Although any phylogenetic tree can reasonably be represented by a
directed acyclic graph, the Phylo module does not attempt to provide a
generally usable graph library – only the minimum functionality to
represent phylogenetic trees. Instead, it provides functions for
exporting Tree objects to the standard graph representations, adjacency
list (dict) and adjacency matrix, using third-party libraries.
to_networkx returns the given tree as a
NetworkX LabeledDiGraph or LabeledGraph
object (depending on whether the tree is rooted). You’ll probably need
to import NetworkX directly for subsequent operations on the graph
object. From this point you can also try using one of NetworkX’s drawing
functions to display the tree, and for simple, fully labeled trees it
may even work – but you’ll have better results with Phylo’s own
draw_graphviz function, discussed above.
import networkx, pylab
tree = Phylo.read("example.xml", "phyloxml")
net = Phylo.to_networkx(tree)
networkx.draw(net)
pylab.show()
Recipes for exporting to other libraries, including ape (via Rpy2) and PyCogent, are available on the Phylo cookbook page.
Many new features for building and processing phylogenetic trees were developed by Yanbo Ye for Google Summer of Code 2013. They are available on the development branch (see SourceCode) but are not yet included in an official Biopython release version. Note that the behavior and API of these features may change before the upcoming official release.
In addition to wrappers of tree construction programs (PHYLIP programs
through EMBOSS wrappers in Bio.Emboss.Applications), now Biopython also
provides several tree construction algorithm implementations in pure
python in the Bio.Phylo.TreeConstruction module.
All algorithms are designed as worker subclasses of a base class
TreeConstructor. All constructors have the same method build_tree
that accept a MultipleSeqAlignment object to construct the tree.
Currently there are two types of tree constructors:
DistanceTreeConstructor and ParsimonyTreeConstructor.
The DistanceTreeConstructor has two algorithms: UPGMA (Unweighted Pair
Group Method with Arithmetic Mean) and NJ (Neighbor Joining).
Both algorithms construct trees based on a distance matrix. So before
using these algorithms, let me introduce the DistanceCalculator to
generate the distance matrix from a MultipleSeqAlignment object. The
following code shows a common way to do this:
>>> from Bio.Phylo.TreeConstruction import DistanceCalculator
>>> from Bio import AlignIO
>>> aln = AlignIO.read('Tests/TreeConstruction/msa.phy', 'phylip')
>>> print(aln)
Alignment with 5 rows and 13 columns
AACGTGGCCACAT Alpha
AAGGTCGCCACAC Beta
GAGATTTCCGCCT Delta
GAGATCTCCGCCC Epsilon
CAGTTCGCCACAA Gamma
>>> calculator = DistanceCalculator('identity')
>>> dm = calculator.get_distance(aln)
>>> dm
DistanceMatrix(names=['Alpha', 'Beta', 'Gamma', 'Delta', 'Epsilon'], matrix=[[0], [0.23076923076923073, 0], [0.3846153846153846, 0.23076923076923073, 0], [0.5384615384615384, 0.5384615384615384, 0.5384615384615384, 0], [0.6153846153846154, 0.3846153846153846, 0.46153846153846156, 0.15384615384615385, 0]])
>>> print(dm)
Alpha 0
Beta 0.230769230769 0
Gamma 0.384615384615 0.230769230769 0
Delta 0.538461538462 0.538461538462 0.538461538462 0
Epsilon 0.615384615385 0.384615384615 0.461538461538 0.153846153846 0
Alpha Beta Gamma Delta Epsilon
As you see, we create a DistanceCalculator object with a string
‘identity’, which is the name of the model (scoring matrix) to calculate
the distance. The ‘identity’ model is the default one and can be used
both for DNA and protein sequences. To check available models for DNA,
protein or all, use the attribute of the calculator dna_models,
protein_models, models respectively.
After the calculator is created with the model, simply use the
get_distance() method to get the distance matrix of a given alignment
object. Then you will get a DistanceMatrix object, a subclass of
Matrix(we will talk about this later).
Now, let’s get back to the DistanceTreeConstructor. We can pass the
DistanceCalculator object and a string parameter(‘nj’ or ‘upgma’) to
initialize it, and then call its build_tree() as mentioned before.
>>> from TreeConstruction import DistanceTreeConstructor
>>> constructor = DistanceTreeConstructor(calculator, 'nj')
>>> tree = constructor.build_tree(aln)
>>> print(tree)
Tree(rooted=False)
Clade(branch_length=0, name='Inner3')
Clade(branch_length=0.182692307692, name='Alpha')
Clade(branch_length=0.0480769230769, name='Beta')
Clade(branch_length=0.0480769230769, name='Inner2')
Clade(branch_length=0.278846153846, name='Inner1')
Clade(branch_length=0.0512820512821, name='Epsilon')
Clade(branch_length=0.102564102564, name='Delta')
Clade(branch_length=0.144230769231, name='Gamma')
While sometimes you might want to use your own DistanceMatrix directly
instead of the raw alignment, we provide another direct way to use both
algorithms.
>>> from TreeConstruction import DistanceTreeConstructor
>>> constructor = DistanceTreeConstructor()
>>> tree = constructor.nj(dm)
>>> print(tree)
Tree(rooted=False)
Clade(branch_length=0, name='Inner3')
Clade(branch_length=0.182692307692, name='Alpha')
Clade(branch_length=0.0480769230769, name='Beta')
Clade(branch_length=0.0480769230769, name='Inner2')
Clade(branch_length=0.278846153846, name='Inner1')
Clade(branch_length=0.0512820512821, name='Epsilon')
Clade(branch_length=0.102564102564, name='Delta')
Clade(branch_length=0.144230769231, name='Gamma')
>>> tree = constructor.upgma(dm)
>>> print(tree)
Tree(rooted=True)
Clade(branch_length=0, name='Inner4')
Clade(branch_length=0.1875, name='Inner1')
Clade(branch_length=0.0769230769231, name='Epsilon')
Clade(branch_length=0.0769230769231, name='Delta')
Clade(branch_length=0.110576923077, name='Inner3')
Clade(branch_length=0.0384615384615, name='Inner2')
Clade(branch_length=0.115384615385, name='Gamma')
Clade(branch_length=0.115384615385, name='Beta')
Clade(branch_length=0.153846153846, name='Alpha')
Unlike DistanceTreeConstructor, the concrete algorithm of
ParsimonyTreeConstructor is delegated to two different worker classes:
the ParsimonyScorer to calculate the parsimony score of a target tree
by the given alignment, and the TreeSearcher to search the best tree
that minimize the parsimony score. A typical usage example can be as
follows:
>>> from Bio import AlignIO
>>> from TreeConstruction import *
>>> aln = AlignIO.read(open('Tests/TreeConstruction/msa.phy'), 'phylip')
>>> starting_tree = Phylo.read('Tests/TreeConstruction/nj.tre', 'newick')
>>> scorer = ParsimonyScorer()
>>> searcher = NNITreeSearcher(scorer)
>>> constructor = ParsimonyTreeConstructor(searcher, starting_tree)
>>> pars_tree = constructor.build_tree(aln)
>>> print(pars_tree)
Tree(weight=1.0, rooted=True)
Clade(branch_length=0.0)
Clade(branch_length=0.197335, name='Inner1')
Clade(branch_length=0.13691, name='Delta')
Clade(branch_length=0.08531, name='Epsilon')
Clade(branch_length=0.041935, name='Inner2')
Clade(branch_length=0.01421, name='Inner3')
Clade(branch_length=0.17523, name='Gamma')
Clade(branch_length=0.07477, name='Beta')
Clade(branch_length=0.29231, name='Alpha')
The ParsimonyScorer is a combination of the Fitch algorithm and
Sankoff algorithm. It will work as Fitch algorithm by default if no
parameter is provide, and work as Sankoff algorithm if a parsimony
scoring matrix (a Matrix object) is given.
Then the scorer is passed to a TreeSearcher to tell it how to evaluate
different trees during searching. Currently, only one searcher
NNITreeSearcher, the Nearest Neighbor Interchange (NNI) algorithm, is
implemented.
By passing the searcher and a starting tree to the
ParsimonyTreeConstructor, we finally get the instance of it. If no
starting tree is provided, a simple upgma tree will be created instead,
with the ‘identity’ model. To use this parsimony constructor, just
simply call the build_tree method with an alignment.
Same as tree construction algorithms, three consensus tree
algorithms(Strict, Majority Rule and Adam Consensus) in pure python are
also implemented in the Bio.Phylo.Consensus module. You can directly
call strict_consensus, majority_consensus and adam_consensus to
use these algorithms with a list of trees as the input.
from Bio import Phylo
from Bio.Phylo.Consensus import *
trees = list(Phylo.parse("Tests/TreeConstruction/trees.tre", "newick"))
strict_tree = strict_consensus(trees)
majority_tree = majority_consensus(trees, 0.5)
adam_tree = adam_consensus(trees)
Instead of using 50% as the cutoff, the majority_consensus method
allows you to set your own one by providing an extra cutoff
parameter(0~1, 0 by default). That means it can also work as strict
consensus algorithm when the cutoff equals 1. One more thing different
to strict and adam consensus tree, the result majority rule consensus
tree has branch support value that are automatically assigned during
calculation.
To get the consensus tree, we must construct a list of bootstrap
replicate trees. So in the Bio.Phylo.Consensus module, we also provide
several useful bootstrap methods to achieve this.
from Bio import Phylo
from Bio.Phylo.Consensus import *
msa = AlignIO.read("Tests/TreeConstruction/msa.phy", "phylip")
msas = bootstrap(msa, 100)
As you see, the bootstrap method accepts a MultipleSeqAlignment
object and generates its bootstrap replicate 100 times. The you can use
them to build replicate trees. While, we also provide a convenient
method to do this.
calculator = DistanceCalculator("blosum62")
constructor = DistanceTreeConstructor(calculator)
trees = bootstrap_trees(msa, 100, constructor)
This time we pass an extra DistanceTreeConstructor object to a
bootstrap_trees method and finally got the replicate trees. Note that
both bootstrap and bootstrap_trees are generator functions. You need
to use list() function to turn the result into a list of alignment or
trees.
Another useful function is bootstrap_consensus. By passing the
consensus method as another extra parameter, we can directly get the
consensus tree.
consensus_tree = bootstrap_consensus(msa, 100, constructor, majority_consensus)
To get the branch support of a specific tree, we can use the
get_support method.
from Bio import Phylo
from Bio.Phylo.Consensus import *
trees = list(Phylo.parse("Tests/TreeConstruction/trees.tre", "newick"))
target_tree = trees[0]
support_tree = get_support(target_tree, trees)
In the above code, we use the first tree as the target tree that we want
to calculate its branch support. The get_support method accepts the
target tree and a list of trees, and returns a tree, with all internal
clades assigned with branch support values.
There are some other classes in both TreeConstruction and Consensus
module that are used in those algorithms. They may not be used commonly,
but might be useful to you in some cases.
The _Matrix class in the TreeConstruction module is the super class
of _DistanceMatrix. They are both actually constructed by a list of
names and a nested list of numbers in lower triangular matrix format.
The only difference between them is that the diagonal elements in
_DistanceMatrix will all be 0 no matter what values are assigned.
To create a _Matrix object:
>>> from Bio.Phylo.TreeConstruction import _Matrix
>>> names = ['Alpha', 'Beta', 'Gamma', 'Delta']
>>> matrix = [[0], [1, 0], [2, 3, 0], [4, 5, 6, 0]]
>>> m = _Matrix(names, matrix)
>>> m
_Matrix(names=['Alpha', 'Beta', 'Gamma', 'Delta'], matrix=[[0], [1, 0], [2, 3, 0], [4, 5, 6, 0]])
>>> print(m)
Alpha 0
Beta 1 0
Gamma 2 3 0
Delta 4 5 6 0
Alpha Beta Gamma Delta
You can use two indices to get or assign an element in the matrix, and the indices are exchangeable.
>>> m[1,2]
3
>>> m[2,1]
3
>>> m['Beta','Gamma']
3
>>> m['Beta','Gamma'] = 4
>>> m['Beta','Gamma']
4
Further more, you can use one index to get or assign a list of elements related to that index:
>>> m[0]
[0, 1, 2, 4]
>>> m['Alpha']
[0, 1, 2, 4]
>>> m['Alpha'] = [0, 7, 8, 9]
>>> m[0]
[0, 7, 8, 9]
>>> m[0,1]
7
Also you can delete or insert a column&row of elements by index:
>>> m
_Matrix(names=['Alpha', 'Beta', 'Gamma', 'Delta'], matrix=[[0], [7, 0], [8, 4, 0], [9, 5, 6, 0]])
>>> del m['Alpha']
>>> m
_Matrix(names=['Beta', 'Gamma', 'Delta'], matrix=[[0], [4, 0], [5, 6, 0]])
>>> m.insert('Alpha', [0, 7, 8, 9] , 0)
>>> m
_Matrix(names=['Alpha', 'Beta', 'Gamma', 'Delta'], matrix=[[0], [7, 0], [8, 4, 0], [9, 5, 6, 0]])
_BitString is an assistant class used frequently in the algorithms in
the Consensus module. It’s a sub-class of str object that only
accepts two characters (‘0’ and ‘1’), with additional functions for
binary-like manipulation (& | ^ ~). Common usage is as follows:
>>> from Bio.Phylo.Consensus import _BitString
>>> bitstr1 = _BitString('11111')
>>> bitstr2 = _BitString('11100')
>>> bitstr3 = _BitString('01101')
>>> bitstr1
_BitString('11111')
>>> bitstr2 & bitstr3
_BitString('01100')
>>> bitstr2 | bitstr3
_BitString('11101')
>>> bitstr2 ^ bitstr3
_BitString('10001')
>>> bitstr2.index_one()
[0, 1, 2]
>>> bitstr3.index_one()
[1, 2, 4]
>>> bitstr3.index_zero()
[0, 3]
>>> bitstr1.contains(bitstr2)
True
>>> bitstr2.contains(bitstr3)
False
>>> bitstr2.independent(bitstr3)
False
>>> bitstr2.independent(bitstr4)
True
>>> bitstr1.iscompatible(bitstr2)
True
>>> bitstr2.iscompatible(bitstr3)
False
>>> bitstr2.iscompatible(bitstr4)
True
In consensus and branch support algorithms, it is used to count and
store the clades in multiple trees. During counting, the clades will be
considered the same if their terminals (in terms of name attribute) are
the same.
For example, let’s say two trees are provided as below to search their strict consensus tree:
tree1: (((A, B), C),(D, E))
tree2: ((A, (B, C)),(D, E))
For both trees, a _BitString object ‘11111’ will represent their root
clade. Each ‘1’ stands for the terminal clade in the list [A, B, C, D,
E] (the order might not be the same, it’s determined by the
get_terminal method of the first tree provided). For the clade ((A,
B), C) in tree1 and (A, (B, C)) in tree2, they both can be represented
by ‘11100’. Similarly, ‘11000’ represents clade (A, B) in tree1, ‘01100’
represents clade (B, C) in tree2, and ‘00011’ represents clade (D, E) in
both trees.
So, with the _count_clades function in this module, finally we can get
the clade counts and their _BitString representation as follows (the
root and terminals are omitted):
clade _BitString count
ABC '11100' 2
DE '00011' 2
AB '11000' 1
BC '01100' 1
To get the _BitString representation of a clade, we can use the
following code snippet:
# suppose we are provided with a tree list, the first thing
# to do is to get all the terminal names in the first tree
term_names = [term.name for term in trees[0].get_terminals()]
# for a specific clade in any of the tree, also get its terminal names
clade_term_names = [term.name for term in clade.get_terminals()]
# then create a boolean list
boolvals = [name in clade_term_names for name in term_names]
# create the string version and pass it to _BitString
bitstr = _BitString("".join(map(str, map(int, boolvals))))
To convert back:
# get all the terminal clades of the first tree
terms = [term for term in trees[0].get_terminals()]
# get the index of terminal clades in bitstr
index_list = bitstr.index_one()
# get all terminal clades by index
clade_terms = [terms[i] for i in index_list]
# create a new calde and append all the terminal clades
new_clade = BaseTree.Clade()
new_clade.clades.extend(clade_terms)
This is how the _BitString is used in the consensus and branch support
algorithms. And I think it can be used in many other conditions.
For example, we can even use it to check whether the structures of two trees are the same.
# store and return all _BitStrings
def _bitstrs(tree):
bitstrs = set()
term_names = [term.name for term in tree.get_terminals()]
term_names.sort()
for clade in tree.get_nonterminals():
clade_term_names = [term.name for term in clade.get_terminals()]
boolvals = [name in clade_term_names for name in term_names]
bitstr = _BitString("".join(map(str, map(int, boolvals))))
bitstrs.add(bitstr)
return bitstrs
def compare(tree1, tree2):
term_names1 = [term.name for term in tree1.get_terminals()]
term_names2 = [term.name for term in tree2.get_terminals()]
# false if terminals are not the same
if set(term_names1) != set(term_names2):
return False
# true if _BitStrings are the same
if _bitstrs(tree1) == _bitstrs(tree2):
return True
else:
return False
To use it:
>>> tree1 = Phylo.read('Tests/TreeConstruction/upgma.tre', 'newick')
>>> tree2 = Phylo.read('Tests/TreeConstruction/nj.tre', 'newick')
>>> tree3 = Phylo.read('Tests/TreeConstruction/pars1.tre', 'newick')
>>> compare(tree1, tree2)
False
>>> compare(tree1, tree3)
True
>>> compare(tree2, tree3)
False
See the Biopython Tutorial sections on sequence alignment and BLAST for explanations of the first few steps shown here.
Search for homologs of a protein sequence using BLAST.
from Bio.Blast import NBCIStandalone, NCBIXML
query_fname = "AAG35789.fasta"
result_handle, error_handle = NCBIStandalone.blastall(
"/usr/bin/blastall", "blastp", "/db/fasta/swissprot", query_fname
)
blast_record = NCBIXML.read(result_handle) # This takes some time to run
Extract the best hits from the BLAST result.
from Bio import SeqIO
from Bio.Seq import Seq
from Bio.SeqRecord import SeqRecord
def get_seqrecs(alignments, threshold):
for aln in alignments:
for hsp in aln.hsps:
if hsp.expect < threshold:
yield SeqRecord(Seq(hsp.sbjct), id=aln.accession)
break
best_seqs = get_seqrecs(blast_record.alignments, 1e-90)
SeqIO.write(best_seqs, "egfr-family.fasta", "fasta")
To help with annotating to your tree later, pick a lookup key here (e.g. accession number) and build a dictionary mapping that key to any additional data you can glean from the BLAST output, such as taxonomy and GI numbers. In this example, we’re only keeping the original sequence and accession number.
Re-align the sequences using Muscle. The program creates an alignment file in Clustal format, “egfr-family.aln”.
from Bio.Align.Applications import MuscleCommandline
cmdline = MuscleCommandline(input="egfr-family.fasta", out="efgr-family.aln", clw=True)
cmdline()
Infer a gene tree using PhyML. First, convert the alignment from step 3 to “relaxed Phylip” format (new in Biopython 1.58):
from Bio import AlignIO
AlignIO.convert("egfr-family.aln", "clustal", "egfr-family.phy", "phylip-relaxed")
Feed the alignment to PhyML using the command line wrapper:
from Bio.Phylo.Applications import PhymlCommandline
cmdline = PhymlCommandline(
input="egfr-family.phy", datatype="aa", model="WAG", alpha="e", bootstrap=100
)
out_log, err_log = cmdline()
Load the gene tree with Phylo, and take a quick look at the topology. (PhyML writes the tree to a file named after the input file plus “_phyml_tree.txt”.)
from Bio import Phylo
egfr_tree = Phylo.read("egfr-family.phy_phyml_tree.txt", "newick")
Phylo.draw_ascii(egfr_tree)
Add accession numbers and sequences to the tree – now we’re using PhyloXML’s extra features.
from Bio.Phylo import PhyloXML
# Promote the basic tree to PhyloXML
egfr_phy = egfr_tree.as_phyloxml()
# Make a lookup table for sequences
lookup = dict((rec.id, str(rec.seq)) for rec in best_seqs)
for clade in egfr_phy.get_terminals():
key = clade.name
accession = PhyloXML.Accession(key, "NCBI")
mol_seq = PhyloXML.MolSeq(lookup[key], is_aligned=True)
sequence = PhyloXML.Sequence(type="aa", accession=accession, mol_seq=mol_seq)
clade.sequences.append(sequence)
# Save the annotated phyloXML file
Phylo.write(egfr_phy, "egfr-family-annotated.xml", "phyloxml")